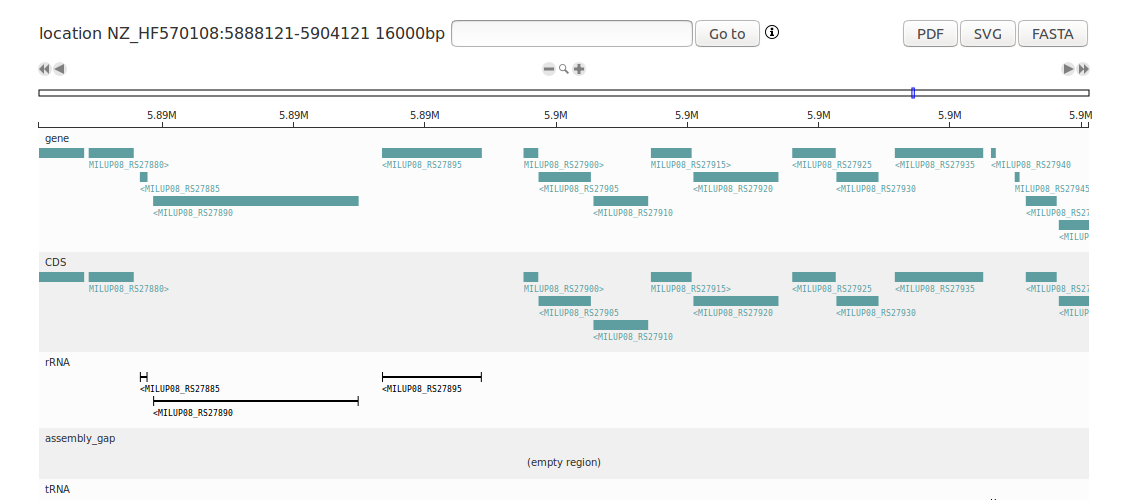
This tutorial shows how to create a D3GB genome browser from a GenBank file with the Python module. This will create a complete genome browser of Micromonospora lupini. Install the D3GB module and write the following code in Python 3:
import D3GB, urllib.request
# Download GenBank file
gbk, headers = urllib.request.urlretrieve('ftp://ftp.ncbi.nlm.nih.gov/genomes/all/GCF_000297395.2_ASM29739v2/GCF_000297395.2_ASM29739v2_genomic.gbff.gz')
# Genome browser generation.
# It creates a genome browser ready to be viewed in Firefox.
# For a server version, ready to be shared with Apache as a Website, set the parameter server=True
gb = D3GB.gbk2genomebrowser(gbk,directory='Micromonospora_Lupini_gbk')
view result